TopAnat: GO-like enrichment of anatomy from expression patterns

TopAnat is a tool to identify and visualize enriched anatomical terms, from the expression patterns of a list of genes.
It allows to discover where genes from a set are preferentially expressed, as compared to a background, represented by default by all expression data in Bgee for the requested species. It is is similar to a Gene Ontology enrichment test, except that it analyzes the anatomical structures where genes are expressed, rather than their GO functional annotations.
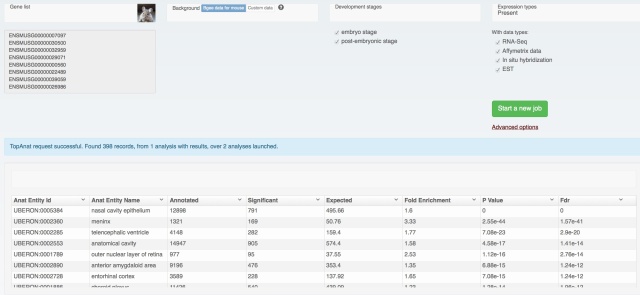
TopAnat uses the classical GO-enrichment approach, comparing terms associated to your gene list to those associated to a background list, and reports terms which are over-represented. The main differences with GO-enrichment are that:
- the ontology used is Uberon rather than GO.
- the association between genes and ontology terms is obtained from gene expression patterns from Bgee rather than GO functional annotations
Like for GO-enrichment, you can use a default background of all genes in your organism with expression data in Bgee, or you can upload your own background set.
- Read the full post
- Try the tool: http://bgee.org/?page=top_anat
November 25, 2015
|
Tweet
Comments Section
Feel free to comment on the post but keep it clean and on topic.
comments powered by Disqus